UC San Diego Team Aims to Broaden Researcher Access to Protein Simulation
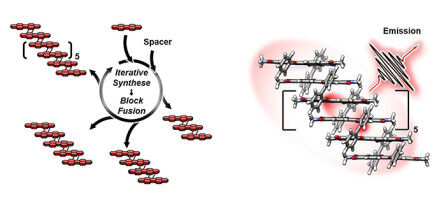
Researchers Enable Unprecedented Sampling Using Desktop Computers with Off-the-Shelf Upgrade. Using just an upgraded desktop computer equipped with a relatively inexpensive graphics processing card, a team of computer scientists and biochemists at the University of California, San Diego, has developed advanced GPU accelerated software and demonstrated for the first time that this approach can sample biological events that occur on the millisecond timescale. These results have the potential to bring millisecond scale sampling, now available only on a multi-million dollar supercomputer, to all researchers, and could significantly impact the study of protein dynamics with key implications for improved drug and biocatalyst development. With some innovative coding, a GPU (graphics processing unit) that retails for about $500, and the widely used software package of molecular simulations called Amber (Assisted Model Building with Energy Refinement), the researchers were able to run a simulation showing the same five long-lived structural states of a specific protein as observed in a simulation conducted by D.E. Shaw Research's Anton , a purpose-built molecular dynamics (MD) supercomputer. The Anton simulation was conducted over a period of slightly more than one millisecond - or 100 times longer than the previous record.