A bacterial enzyme could be the key to sustainable production of the plastic building block ethylene
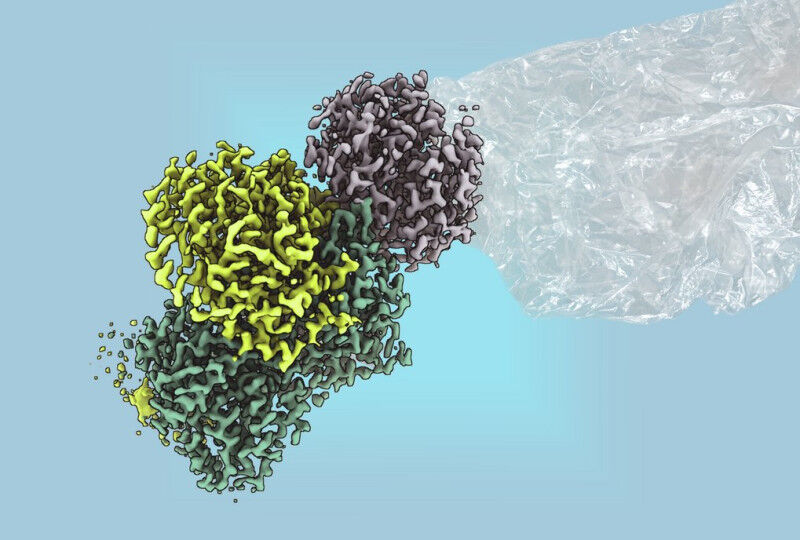
- The conventional production of ethylene releases large quantities of greenhouse gases. A bacterial enzyme from Rhodospirillum rubrum could make it possible in future to produce the plastic building block ethylene without CO2 emissions.
- Researchers at the Max Planck Institute for Terrestrial Microbiology have investigated the structure and function of the enzyme.
- The results provide the biochemical basis for developing a more sustainable biotechnological production of ethylene - and at the same time shed light on early biochemical processes on Earth.
Ethylene is currently produced on a large scale from fossil fuels to meet the current demand for plastics and chemical base materials. This necessitates the search for new, renewable processes. The use of bacterial metabolic pathways could be a solution in the future, but only a few enzymes are known in nature that can produce ethylene. These often require energy-rich substrates. In addition, they also release CO2 as a by-product.
The scientific community therefore reacted with great interest when the enzyme methylthioalkane reductase was discovered in the bacterium Rhodospirillum rubrum a few years ago. With its help, the bacterium produces ethylene under oxygen-free conditions without producing CO2.
Very special enzymes: the -Great Clusters of Biology-
The fact that production takes place under oxygen-free conditions is also a problem: due to the considerable challenges involved in purifying and handling the oxygen-sensitive metalloenzymes, methylthioalkane reductase could previously only be studied in cell cultures. Direct proof of its activity outside the cell has been lacking to date. Many important questions remain unanswered before a possible biotechnological use: How does the catalytic mechanism work and what properties determine it?
Researchers at the Max Planck Institute for Terrestrial Microbiology in Marburg, led by Johannes Rebelein, have now purified the enzyme and elucidated its structure in a collaboration with RPTU Kaiserslautern. The researchers made an exciting discovery: -The reaction is driven by large, complex iron-sulphur clusters, which were previously thought to occur only in nitrogenases, the oldest enzymes on earth-, explains Ana Lago-Maciel, PhD student and first author of the study. -The enzyme is the first non-nitrogenase enzyme known to contain these metal clusters
Nitrogenases evolved billions of years ago. They are the only known enzymes that make gaseous nitrogen from the atmosphere available for life by reducing nitrogen to enable its incorporation into biomolecules such as DNA and proteins. This unique achievement is based on a special feature: large and complex iron-sulphur clusters. Due to their structural complexity and geochemical importance, the metal clusters of nitrogenases belong to the so-called -Great Clusters of Biology-.
The work also reveals the biochemical and structural background of a geochemically important source of hydrocarbons. -In fact, the enzyme is remarkably versatile," explains Johannes Rebelein, head of the research project. -It can produce a range of hydrocarbons, including ethylene, ethane and methane
Blueprints for a more sustainable plastics industry
The very different substrate spectrum of the enzyme compared to nitrogenases opens new doors for understanding how the protein structure determines the reactivity of the metal clusters. -Our work provides the basis for biotechnologically taming these enzymes and adapting their product spectrum to our needs," says Johannes Rebelein.
In addition, the results are an indication of the evolution and role of the -Great Clustersin the past. -The results indicate that structurally similar enzymes were already using these clusters for reductive catalysis long before nitrogenases themselves evolved. This is a dramatic shift in our understanding of this crucial part of the history of life on Earth


