Adhesion GPCRs are a group of cell surface sensors that are associated with many bodily functions and diseases. However, they have not yet been sufficiently investigated in order to use them for therapies. The Collaborative Research Center 1423 at Leipzig University aims to change this. Scientists from the Faculty of Medicine have now developed an innovative numbering system for the GAIN domain, a protein domain that is common to all adhesion GPCRs. It should help to better classify the role of this protein domain in diseases and pave the way for more precise experimental approaches. The work was published in journal Nature Communications.
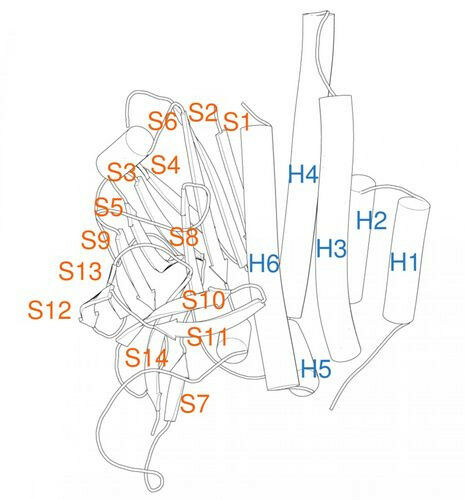
Adhesion G protein-coupled receptors form a large class of membrane proteins that recognize chemical and mechanical stimuli in the body. The GAIN domain plays a central role in the activation of adhesion GPCRs. Until now, research has been limited by the low similarity of the amino acid sequence of different GAIN domains, which has hampered knowledge transfer and comparative analyses. The new numbering system developed by researchers at Leipzig University now creates a uniform basis for the precise transfer of research results between different model systems and humans. It is based on the structural analysis of over 14,000 modeled structures of the GAIN domain, which were generated using the latest AI methods.
Peter Hildebrand, Professor of Membrane and Cell Biophysics at the University of Leipzig, who led the study with international participation, says: "Our new numbering system is an important step in GPCR research. It facilitates basic research and promotes comparative studies. It provides a solid basis for further biochemical, bioinformatic and structural biology research." The newly developed system makes it possible, for example, to better understand disease-relevant mutations in GAIN domains, which provides a deeper insight into their role in diseases such as cancer.
Analysis can be used for kidney disease
In the current publication, the scientists found that their numbering system also enables the analysis and comparison of the GAIN domains of polycystic kidney disease proteins. This genetic disease leads to the formation of fluid-filled cysts in the kidneys and other organs.
Florian Seufert, first author of the current publication and doctoral student at the Institute of Medical Physics and Biophysics, says: "The interdisciplinary work in the SFB1423 and our contacts in Berlin and Copenhagen bring a variety of perspectives to a project like this. This has enabled us to develop our system in a user-friendly way, opening the door to numerous new research opportunities. We are already using the system to support two projects in which we are further investigating the function of the GAIN domain."
The new numbering system has been integrated into the internationally widely used database on GPCRs, facilitating global access and knowledge exchange for researchers.
Original publication in Nature Communications: Generic residue numbering of the GAIN domain of adhesion GPCRs. DOI: https://doi.org/10.1038/s41467-024-55466-6
The current publication was produced as part of the Collaborative Research Center 1423 "Structural Dynamics of GPCR Activation and Signal Transduction", which is funded by the German Research Foundation (DFG). Peter Hildebrand is currently also involved in a publication on GPCR research in the journal Nature Reviews Drug Discovery .

