 3D illustration of a coronavirus binding to a human cell.
3D illustration of a coronavirus binding to a human cell.
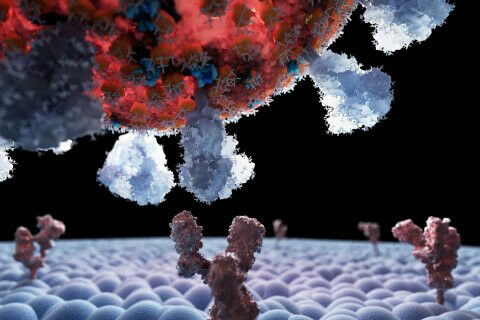
3D illustration of a coronavirus binding to a human cell. (©Christopher Hohmann, LMU Munich) A key factor in the rapid spread of the so-called British coronavirus variant appears to be stronger attachments between the virus and human cells. In a study led by Utrecht University professor Jan Lipfert, scientists show that the variant has a significantly stronger attachment to human cells compared to the original strain. This finding may help explain why the British variant became so dominant during the early stages of the pandemic. As the world grappled with the initial waves of coronavirus infections, mutated versions of the virus soon emerged. A variant initially dubbed the "British variant" quickly replaced the original Covid strain as the most prevalent one. Named after the countries where they were first identified, these "variants-of-concern" sparked questions about what made certain strains spread more than others.
TO READ THIS ARTICLE, CREATE YOUR ACCOUNT
And extend your reading, free of charge and with no commitment.
Your Benefits
- Access to all content
- Receive newsmails for news and jobs
- Post ads


